
By analyzing RNAs that have been believed as "non-coding", we will identify novel proteins that have been overlooked. In this way, we expect to reveal molecular mechanisms of biological phenomena that have not been well explained so far.


By analyzing RNAs that have been believed as "non-coding", we will identify novel proteins that have been overlooked. In this way, we expect to reveal molecular mechanisms of biological phenomena that have not been well explained so far.



PROJECT 1
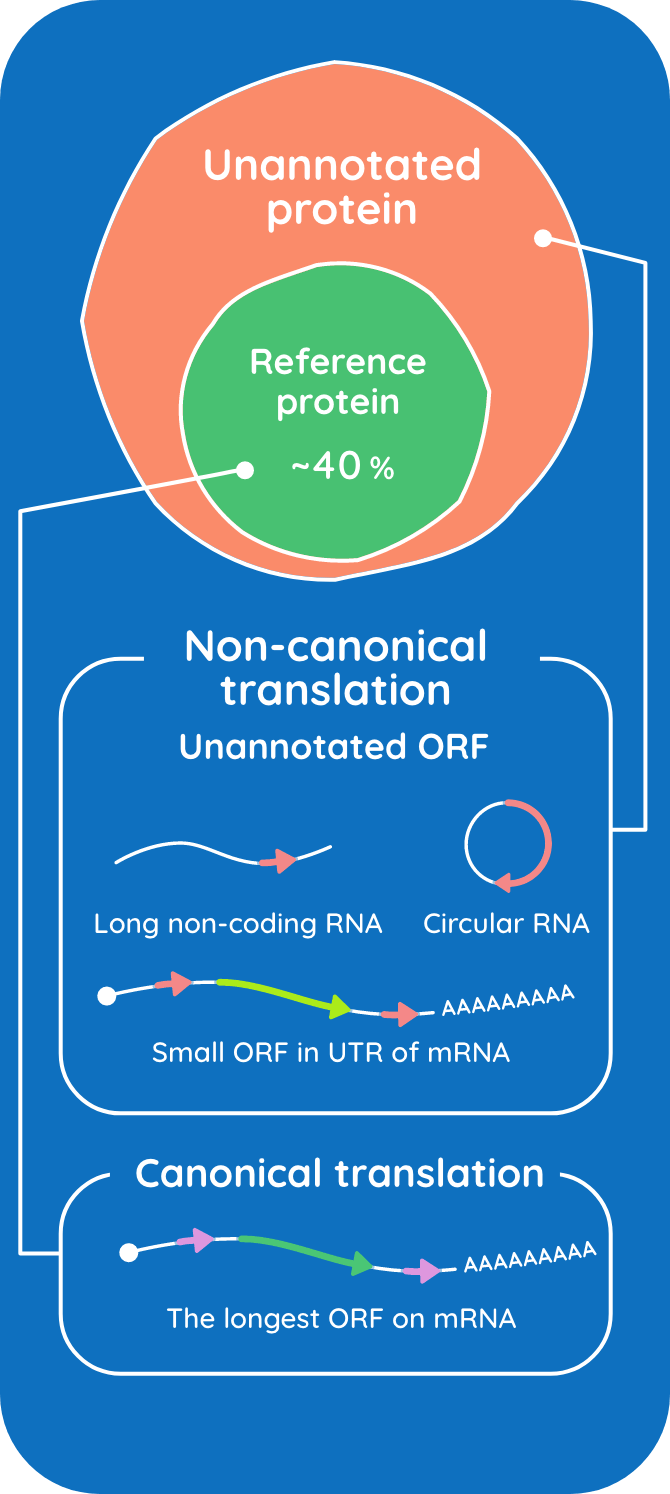
For a long time, it was thought that only the longest open reading frame (ORF) in messenger RNA, the coding sequence (CDS), was translated into protein. However, it has been suggested that a myriad of small ORFs (unannotated ORFs: uaORFs) may also be translated to produce small proteins. We aim to identify and elucidate the roles of such small proteins by using molecular biology and biochemical techniques.
PROJECT 2
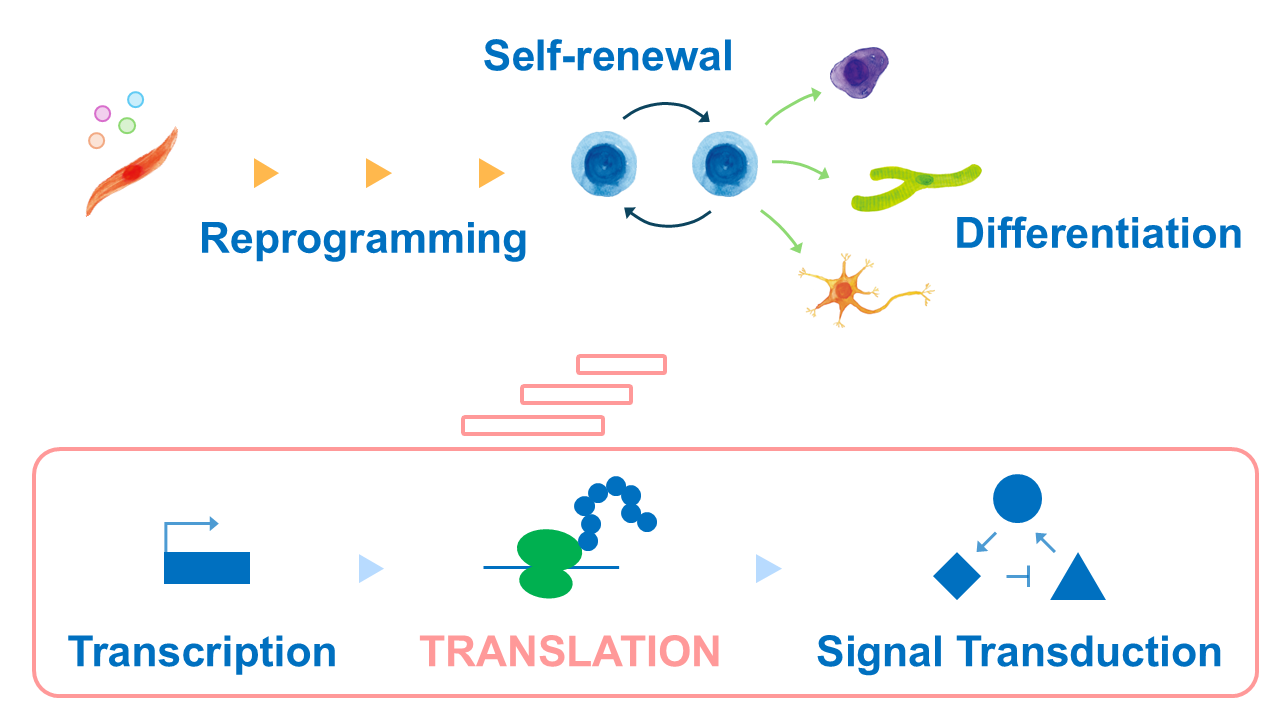
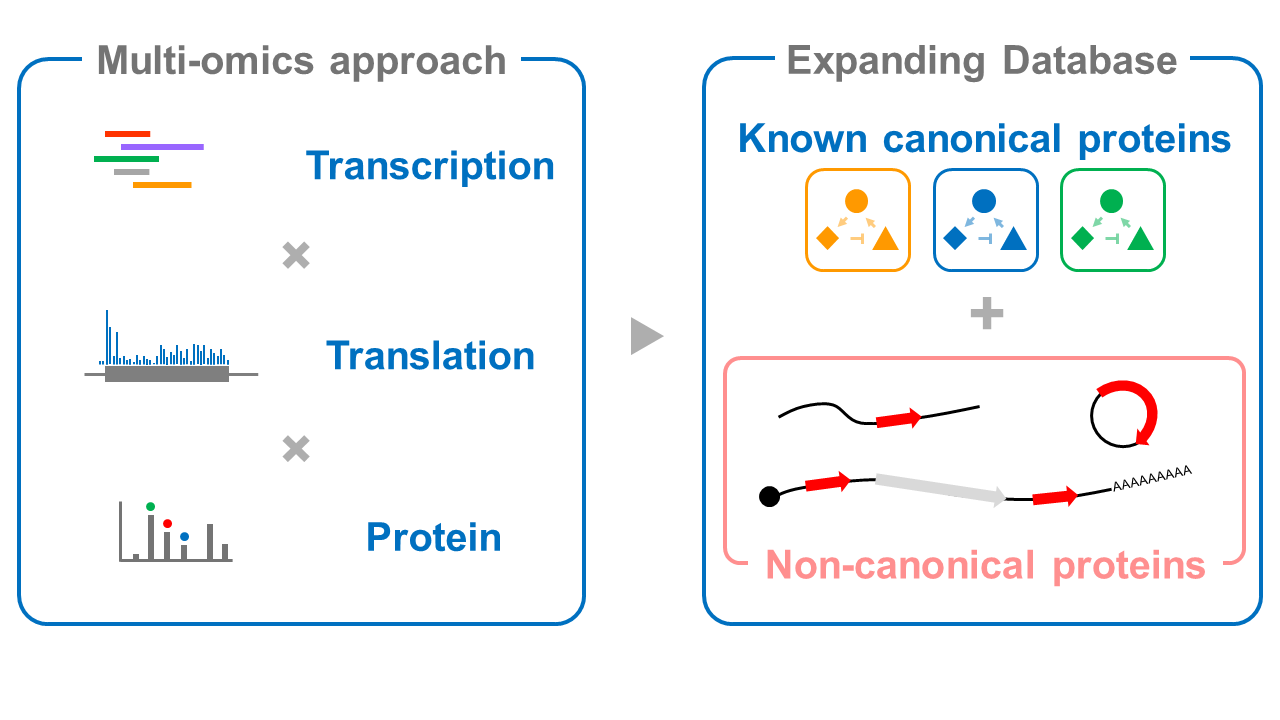
The mechanism of pluripotency has been studied focusing on transcription, which generates mRNA from DNA, and signal transduction, which is a workplace of functional proteins. However, many unknowns still exist, and research from a different angle is needed. Therefore, we aim to gain a deeper understanding of the mechanisms of pluripotency by focusing on translation regulation, which acts as a bridge between transcription and signal transduction.
PROJECT 3
Pluripotent stem cells continue to proliferate without aging. Our goal is to understand the translational control mechanisms that underlie the "forever young" ability to develop technologies that will help cells and individuals live long and healthy lives.